Using networks and supercomputing to dig deeper into multi-omics data
Speakers
Event series
Content navigation
Description
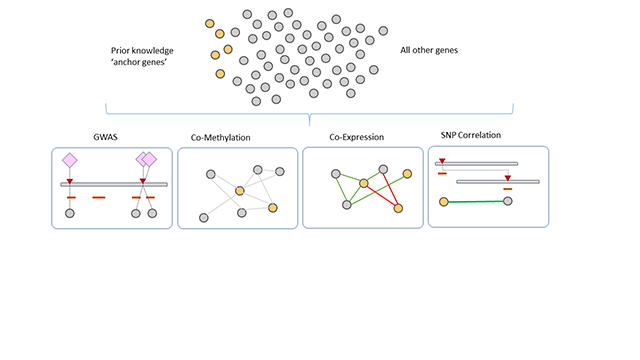
The increasing proliferation of biological assays, high throughput phenotyping and computational prediction has resulted in an enormous wealth of biological data in many different species. These data layers (e.g. genomic, transcriptomic, metabolomic, protein-protein interaction, phenomic) are developed with the goal of understanding the operation of overarching systems and discovering the basis for emergent phenotypes. Each data layer can be interpreted within the context of just that dataset, which often provides useful, but limited, insights. This is because biological elements rarely operate in isolation within and between the cellular environment, so data from a single layer tells only part of the story, or possibly even misleads. It is clear that a multi-layer systems-biology view is necessary, and that the more independent lines of data we have available the more we are able to deconvolute the complexities of the system. But analyzing data of that scale is a huge challenge. A useful approach is to represent data from assays of expression, GWAS, transcription factor (TF) binding, miRNA, protein-protein interactions (PPI) and more as networks. Such networks provide a natural way to visualise and mine vast amounts of data from multiple sources because any biological entity, such as a gene, protein, metabolite or trait, can be represented by a network node, while interactions between entities are represented by edges between the nodes. In this talk I will discuss how one can construct a variety of innovative networks that are only possible with high performance computing methods, and use network analysis methods to gain deep biological insights about relationships within and across multi-omic layers.
Location
Eucalyptus Seminar Room Rm S205, Level 2, RN Robertson Building (46)
