E&E PhD Exit Seminar: The treelikeness assumption: Investigating conflicting signal in the animal tree of life
Phylogenetics is the science of reconstructing the evolutionary history of groups of species or individuals. Most phylogenetic methods include the treelikeness assumption, which states that every site in an alignment shares an identical evolutionary history.
Speakers
Event series
Content navigation
Description
Phylogenetics is the science of reconstructing the evolutionary history of groups of species or individuals. Most phylogenetic methods include the treelikeness assumption, which states that every site in an alignment shares an identical evolutionary history. However, the treelikeness assumption is violated by biological processes (such as introgression, hybridisation, and incomplete lineage sorting) or analytical processes (such as concatenation or alignment error). Although conflicting evolutionary histories have been found within single genes, most phylogenetic protocols do not consider the impacts of this non-treelike data on tree accuracy.
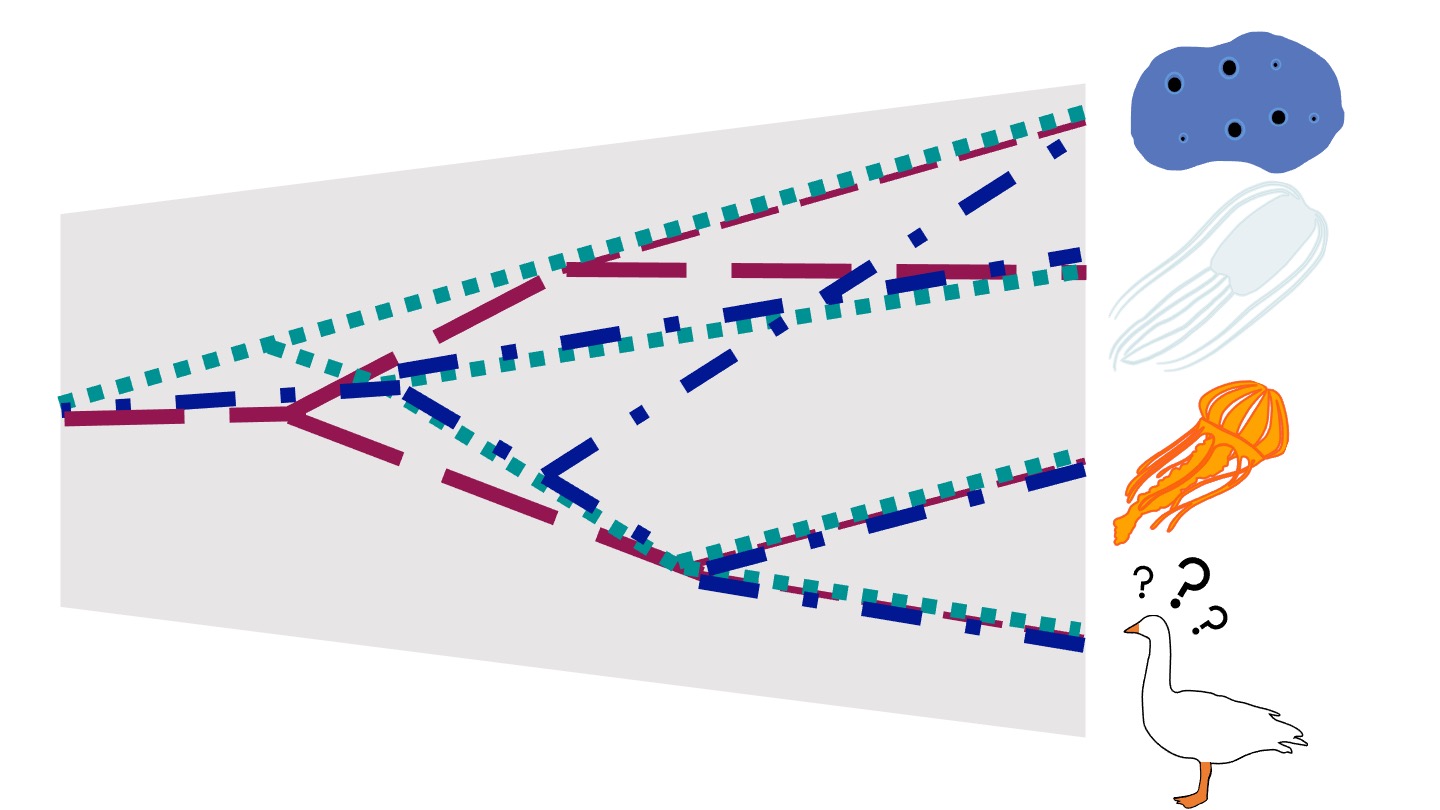
The Metazoan tree of life describes the evolutionary history of all animals and consists of five clades: the sponge clade Porifera, the comb jelly clade Ctenophora, Placozoa, Cnidaria, and Bilateria. Historically, sponges were thought to be the sister to all other Metazoan clades, but recent molecular studies have found evidence for Ctenophora as the sister to all other animals. There is currently no consensus on the relationship between different Metazoan clades. This tree is particularly complex to resolve, due to the deep timescales since divergence events between Metazoan clades (>500 million years), results in distantly related species with highly divergent genomes. In addition, animals underwent a rapid radiation towards the root of the Metazoan tree, resulting in short branches between clades which are difficult to resolve.
In this seminar, I investigate conflicting signal within the Metazoa under a treelikeness framework. First, I present a new metric to quantify treelikeness within any multiple sequence alignment, and present results for the first (to my knowledge) comprehensive benchmarking of metrics for treelikeness from the literature. Secondly, I apply tests for recombination to empirical phylogenetic datasets and quantify the impact of including non-treelike loci on tree accuracy. Thirdly, I relax the treelikeness assumption, allowing a single multiple sequence alignment to have 2 or more underlying evolutionary histories, to determine whether a single tree is sufficient to describe the evolutionary history of the Metazoans. Finally, I interrogate published Metazoan datasets to understand the biological or analytical process that generated conflicting phylogenetic signal.
Location
Please note: this seminar will be held in the Eucalyptus Rm and via Zoom, details are included below.
Eucalyptus Room, Rm S205, Level 2, RN Robertson Building (46)
Please click the link below to join the webinar:
https://anu.zoom.us/j/88156927904?pwd=KzM3QUV6dGJOazRLSXpEMENVUllxQT09
Passcode: 563958
Canberra time: please check your local time & date if you are watching from elsewhere.
